Introduction: AI in Drug Discovery.
AI in drug discovery: AI-enabled drug discovery (AIDD), a sector that has experienced a breakthrough year in funding with $4.1B committed in 2021 alone, is a prime example of platform techBio. AIDD involves the application of computational tools to the drug discovery and development process with the goal of reducing the time and capital required to take a drug from research to masses. As we move forward with AIDD, we hope to move from drug discovery to deliberate drug design.
First AI designed drug in human trials.
In order for a new drug to be discovered, we have to find 1) a biological target for the drug, and 2) a drug that will work on the target. In the grander scheme of things, a target is any cellular structure that affects disease biology in any way. Biological structures include proteins, enzymes, hormones, strings of nucleic acids, and many other kinds of molecules. The reality is that there are literally millions of possible targets for any given disease when we take this into consideration. Additionally, there are maybe another million possible drugs that could bind to the target or otherwise interact with it to cause the desired therapeutic effect. It would be difficult to test all those combinations using brute force in a wet lab.
Source - YouTube | The Age of AI.
To say that early-stage drug discovery is similar to fitting a lock to a key is an understatement. In order to open the door, we must first figure out the lock (target), and then we must determine the key (drug) that will open the door. The only thing that needs to happen before we can accomplish any of this is to decide what room (disease/indication) we want to open (or close) in the first place. In biology, nothing is without layers of complexity. Therefore, there are some indications that have been shown to be extremely difficult to target, while others don’t have obvious targets that can be drugged. On the surface, it appears that many doors are locked from the inside.
Robotic Surgery
The pharmacology research can be divided into two types: the classical approach and the reverse approach
The idea of conventional pharmacology is that you know what the diseased state is and what the healthy state is, so you go out and look for compounds that cause the desired phenotypic change by identifying what is known as a target. In reverse pharmacology, you identify which molecules/targets are involved in the disease and then screen for compounds that bind or inhibit those molecules/targets in some way. Target identification is typically skipped over in favor of experimental validation and hit generation. Regardless, the process is similar since it is aimed at finding a target that can be modulated and a drug candidate to modulate it.
AI test that detects heart disease in just 20 seconds
Simplified Drug Discovery Process
1. Target Identification and Validation
It is important to understand that target identification is concerned with the identification of a few proposed biological sites or molecules that potential drugs can interact with in order to modify disease activity.
In the past, disease targets have been discovered by studying specific pathways that affect the health of patients. Studies of this type are heavily based on biochemical principles and are meant to explain the relevance of the target in the broader context of the disease. In the context of this question, a relevant question to ask is whether inhibiting or activating this target has the ability to change the expression of the disease?
First Robotic Surgery
Even if you find a promising target that affects the disease activity in a specific way, it might not be the one that is best for the job. Because you only get what you screen for, you wouldn’t know there is a better target out there, because you don’t know it exists. The benefit of machine learning here is that screens will not be limited by the experimental capacity of the experiment.
When a target is identified, the focus shifts from identifying it to validating it. An identification of target is concerned with finding a molecular element that has been implicated in a disease of interest, whereas validation is concerned with making sure that modulating such a target will result in the desired effect. Various biochemical experiments can be conducted in order to validate targets, including generating gene knockouts, measuring protein interactions, and evaluating binding and/or kinetics.
How Can RPA Help In Healthcare?
2. Hit Identification and Lead Generation
A valid target must then be followed up by the search for “hits” or molecules that favorably interact with the target, i.e. potential drug candidates. In terms of what type of drug is desired, the process can range from massive small molecule library screening against the target to intentionally engineering molecules (often proteins) that bind to the target.
AIDD companies are starting to develop a preference for deliberate drug design by anticipating properties and structure constraints. Also, some companies screen against panels of already FDA-approved drugs to find new uses for them.
Impact Of Automation In Healthcare
Depending on the researcher, the terms “target”, “hit”, and “lead” can have different meanings. However, the approach to drug discovery remains the same. We will use the term targets in this article to refer to biological sites, hits to represent potential drug candidates that interact with targets, and leads to represent candidates for further development.
Identification of hits results in the identification of promising drug candidates that can then be further evaluated in later stages of research and ultimately trials. Not only are the target/ligand-binding and disease-affect properties of potential hits being evaluated, but also the pharmacokinetic (PK) and pharmacodynamic (PD) properties of the hits are being assessed at this stage as well.
The Impact of Artificial Intelligence in Ophthalmology
The properties of these compounds are often referred to as ADME/Tox (absorption, distribution, metabolism, elimination, and toxicity) and can be tested via both in silico (computer-run studies) and wet-lab laboratories to filter out undesirable drug candidates as initial hits are refined in something called “hit to lead” or lead generation.
It is quick and inexpensive to perform in silico ADME/Tox tests on hits before they are advanced. A good example of such a test may be Schrodinger’s QikProp program. Additionally, Lipinski’s Rule of 5 is still engaged to help with hit screens. Lipinski’s rules were developed by Christopher Lipinski during the late 1990’s as a guide for the design of drugs based on clinical data from Merck and Pfizer regarding drug candidates.
The role of AI in vaccine distribution
Although these rules are highly specific and formulation-dependent, they are astonishingly accurate and underscore the value of analyzing large volumes of data to guide drug discovery and optimization.
These generalizations, even though they are not always adhered to exactly, are useful guides for streamlining the search for relevant results. In the event that no promising hits are discovered for a given target, then it’s time for a fresh start.
Artificial Intelligence in Healthcare Business Process Improvement
3. Lead Optimization
As soon as hits have been identified, the focus turns to lead optimization or advancing the lead candidate to the next level of preclinical development.
A major focus of this conversion is to improve the properties (ADME/Tox, structural, etc.) of the lead in order to increase efficacy and decrease toxicity, characteristics which are in line with the safety & efficacy profiles that the FDA will be looking for before Investigational New Drug approval and trials. Studies using animal models, dosing studies, and in vitro and in vivo assays are performed in order to quantify target/lead interactions as well as general toxicities.
Focus is on improving the drug candidate’s lackluster aspects while maintaining its favorable properties, a challenging task given the ambiguity and interdependence of biological structure and phenotype. It is therefore important to take care when enhancing the absorption profile of a lead that developers are mindful of not also enhancing aggregation. If you can think of this as taking one side of a Rubik’s Cube at a time, then you can imagine you are solving a multidimensional puzzle.
Computing models are extremely useful in situations like this. When the candidate is being developed, it needs to be as close to perfect as possible. There are multiple characteristics being refined in parallel at any one time, which makes it possible to create endless variations of the lead over a period of several years. Algorithms are used to predict and test modifications in a more data driven and efficient manner.
How Far is Too Far? | The Age of A.I. | S1 | E1
There is a great deal of value and complexity locked up in lead optimization, to the point where several companies only focus on it. Upon the emergence of a successful hit and lead combination, the optimized lead candidates move on from the drug discovery phase and begin the longest stage of drug development, clinical research.
Artificial Intelligence in Drug Discovery
This context makes it clear that there is plenty of room for AI to disrupt discovery, not only by accelerating the discovery process, but also by identifying higher-fidelity candidates as well. The use of computational approaches may move the field away from discovery and towards outright design.
Types of ML models in the drug discovery process.
A comprehensive review of all computational methods that can be applied to drug discovery would be like boiling the ocean. Our attempt here is to highlight a few common approaches employed in computational biology, a breakdown borrowed from Professor Manolis Kellis’ lecture series on the topic.
Simulation: a structure-based method for determining molecular structure and dynamics, such as ligand docking and binding profiles with virtual simulations. It is possible to determine the target and potential ligands using this strategy, but it requires sorting through a large number of compounds and checking the dynamics in the wetlab. The use of virtual simulation may help elucidate how a target and ligand interact to guide further experiments but does little to speed up the process of drug discovery since additional experiments are still needed at each stage.
In silico screening: A computer-based version of high throughput screening, in silico screening is the process of searching online libraries for molecular compounds by using algorithms. It requires access to relevant and organized datasets, whether public or proprietary. While virtual screening can be useful for targeted applications, it still has the drawback of only getting what you screen for. If you predict certain properties like solubility and biodistribution, you can screen for the best candidates, but even if a virtual screen returns a few hits, you have no way of knowing whether those are indeed the best hits; in other words, you’re left in a position where the local maxima look deceiving like the global maxima.
New drug design: As we move from drugs-hunting to deliberately engineering optimal drug candidates, this group of methods is the most interesting. In contrast to simulation and virtual screening, which involve mapping the chemical space (known compounds) to the functional space (desired properties), novel drug design utilizes computational techniques to find or design the right chemistry.
Visualization of ligand binding a docking site through PyMOL, a common tool used in Simulation.
Datasets in AI Drug Discovery
Since algorithms utilized by AIDD companies have many similarities (deep learning, computer vision, natural language processing, etc. ), it is often the data used that distinguishes one company from another and adds to its competitive advantage. Almost always, the objective is to find a target and a lead that work together to modulate disease. The two types of data that are most compelling for figuring out this are sequence and compound library data, and imaging data, with each company using both, from proprietary and publicly available data.
The sequence data and compound libraries – are data collections derived from molecules and/or molecular sequences at the DNA, RNA, and protein level. Public databases including UniProt and PDB contain most of this data. It is unlikely that epigenetic sequencing data will cease to be a major consideration when drug discovery becomes more design-forward and data-driven. The Omics data is particularly useful in building these datasets because it incorporates insights from spatial transcriptomics, genomics, proteomics, and multiomics. Phenomic, DeepMind’s AlphaFold, Exscientia, Enveda Bioscience, Unnatural Products, and Valence Discovery are among the companies that rely heavily on sequencing data and compound libraries.
The imaging data—cellular/molecular imaging is one of the best ways to gain insight into how molecules interact and how reactions occur. In addition to x-ray crystallography, cryoEM, and other microscopy platforms, there are tools that can generate detailed information about cellular interactions. As a rule of thumb, companies building platforms around imaging data have a unique advantage over their competitors in building proprietary datasets of cellular/molecular images since this type of data needs to be granular and it isn’t always easy to find public datasets tailored to specific tasks. These companies include Recursion Pharmaceuticals , Eikon Therapeutics , Gandeeva Therapeutics, and AbCellera.
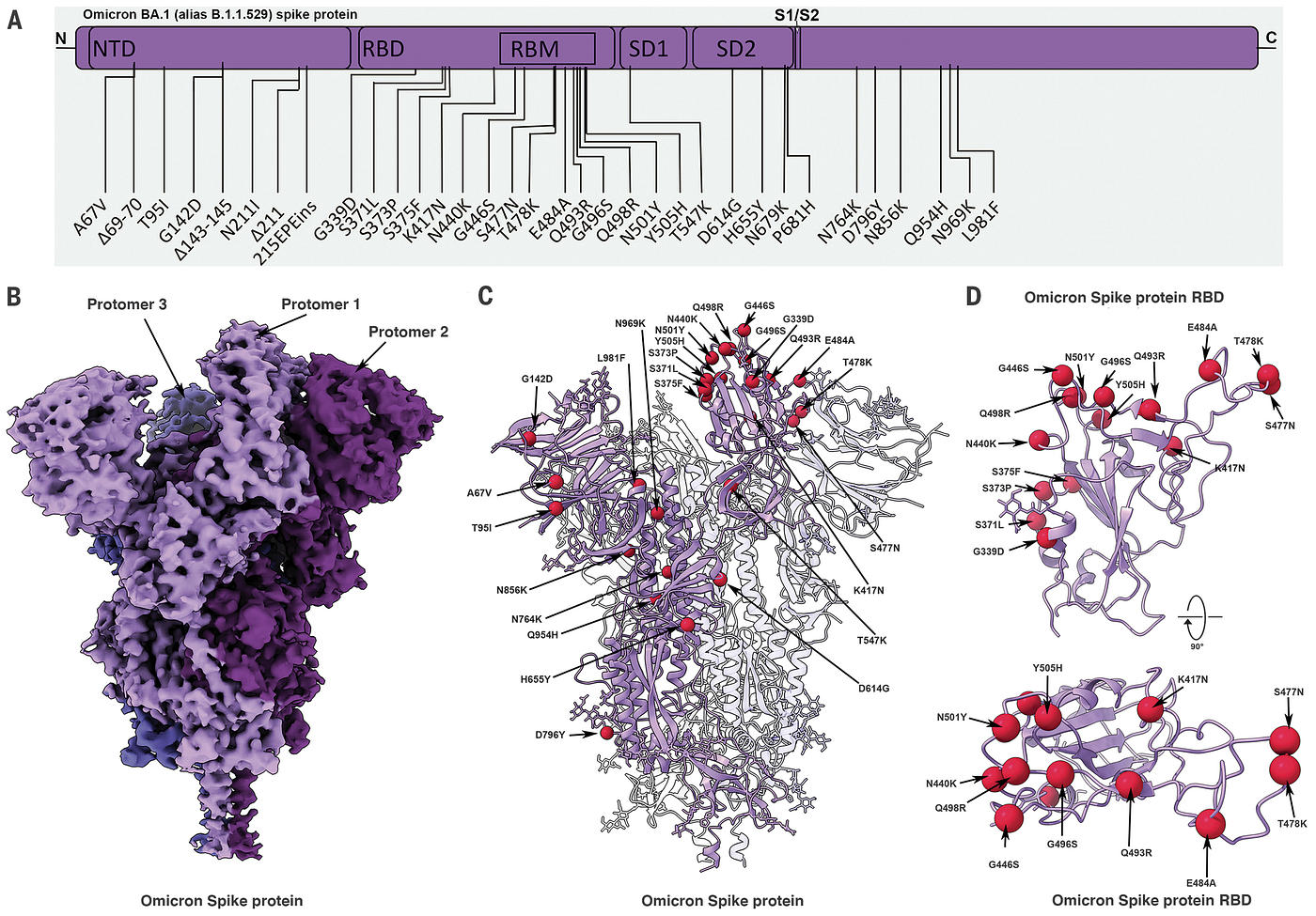
CryoEM data (left) from SARS-CoV-2 Omicron variant.
Artificial Intelligence and Otolaryngology.
Algorithms in AIDD
As a follow up to the data collection, various algorithms are now utilized to make sense of the data and create predictions for finding targets or finding hits, depending on what stage you are in the drug discovery process. Below are three computational methods that are explained in more detail:
Deep Learning – Unstructured data is analyzed using deep learning, which is a subset of machine learning. While deep learning requires a large amount of data, that data does not have to be tagged. The software can be used for a variety of drug discovery tasks, including advanced image analysis, structure prediction, and novel chemical design. It is very common for different algorithms to be implemented into the same platform; for instance, one set of algorithms can be used to identify the binding of target/ligand, while another set optimizes potential drug candidates for ADME/Tox properties, and another ensures that the molecules suggested are not infeasible to synthesize at scale. Gradient descent, reinforcement learning, and neural networks (concurrent, convolutional, graph, etc) are some examples of deep learning frameworks that may be applied to AIDD.
Computer Vision – In computer vision, molecular and cellular images are sorted into numbers that can be processed with greater throughput and used to make accurate predictions. Computer vision is a classic example of drug discovery in Recursion’s technology that analyzes images of diseased vs healthy cells to better understand disease biology and develop new targets and drug candidates.
Natural Language Processing (NLP) — Understanding how nature designs molecules and biology (or “writes” words) and predicting structure accordingly. As an example of NLP applied to drug design, Nabla Bio’s* approach of reading amino acid sequences to predict the structure and function of proteins provides an example of NLP applied to drug design.
Artificial Intelligence in Healthcare.
Machine learning cannot be applied to all aspects of drug discovery-it is common to see companies working across the entire spectrum of drug discovery and design, from mapping cellular interactions with computer vision for improving target discovery to using natural language processing to predict entirely new proteins to using patient data to pinpoint genetic variations. The application of ML to drug discovery can take place at any stage of the process and, ideally, as we gain a better understanding of biological interactions and the limitations of AI, algorithms will increasingly be used to develop the drugs we want outright.
AI-powered drug design companies will easily surpass top-ten pharma companies with regard to capital efficiency to the masses. The first AI-discovered drug has yet to receive the highest level of clinical validation – FDA approval and post-market surveillance – despite our optimism.
How can Artificial Intelligence help with the Coronavirus (Covid-19) vaccine search?
Even though the jury is still out, early data indicates that AI can improve drug discovery by increasing hit rates, increasing efficiency, decreasing the time to clinic, and improving patient outcomes.
